There are over 7,000 unique rare diseases, some of which affecting 3,500 or fewer patients in the US. Due to clinicians’ limited experience with such diseases and the considerable heterogeneity of their clinical presentations, many patients with rare genetic diseases remain undiagnosed. While artificial intelligence has demonstrated success in assisting diagnosis, its success is usually contingent on the availability of large labeled datasets.
Here, we present SHEPHERD, a deep learning approach for multi-faceted rare disease diagnosis. SHEPHERD is guided by existing knowledge of diseases, phenotypes, and genes to learn novel connections between a patient’s clinico-genomic information and phenotype and gene relationships. We train SHEPHERD exclusively on simulated patients and evaluate on a cohort of 465 patients representing 299 diseases (79% of genes and 83% of diseases are represented in only a single patient) in the Undiagnosed Diseases Network. SHEPHERD excels at several diagnostic facets: performing causal gene discovery (causal genes are predicted at rank = 3.52 on average), retrieving “patients-like-me” with the same gene or disease, and providing interpretable characterizations of novel disease presentations.
SHEPHERD demonstrates the potential of artificial intelligence to accelerate the diagnosis of rare disease patients and has implications for the use of deep learning on medical datasets with very few labels.
Rare diseases affect 300-400 million people worldwide, yet each disease has a very low prevalence, affecting no more than 50 per 100,000 individuals. Due to their low prevalence, most front-line clinicians lack experience with the disorders, resulting in many specialty referrals and expensive clinical workups for patients across multiple years and institutions. Furthermore, patients with the same disorder can present variable symptoms, disease severity, and age of onset. These challenges make rare disease diagnosis extremely difficult; approximately 70% of individuals seeking a diagnosis and up to half of the suspected Mendelian conditions remain undiagnosed.
Given a set of phenotypes describing a patient’s clinical presentation, SHEPHERD performs a multi-faceted patient diagnosis to identify causal genes, retrieve “patients-like-me” with the same causal gene or disease, or provide interpretable characterizations of novel disease presentations. SHEPHERD assists in the rare disease diagnosis at multiple points:
- to find similar patients after the patient’s clinical workup,
- to identify candidate genes after genome sequencing analysis or in conjunction with the clinical case review, and
- to characterize the patient’s disease and find other patients with the same causal gene or disease.
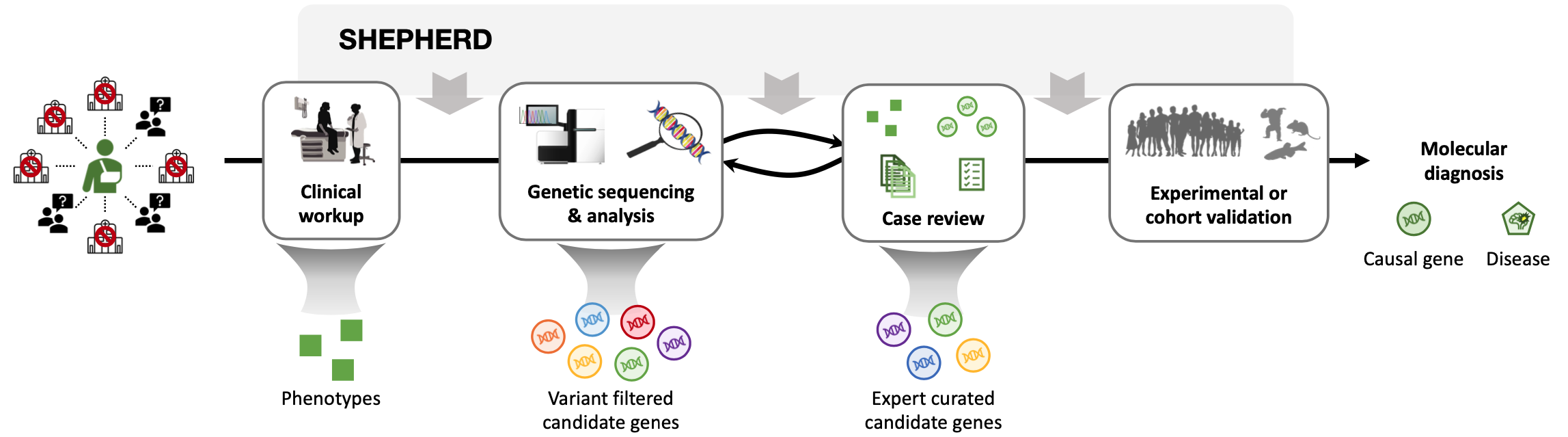
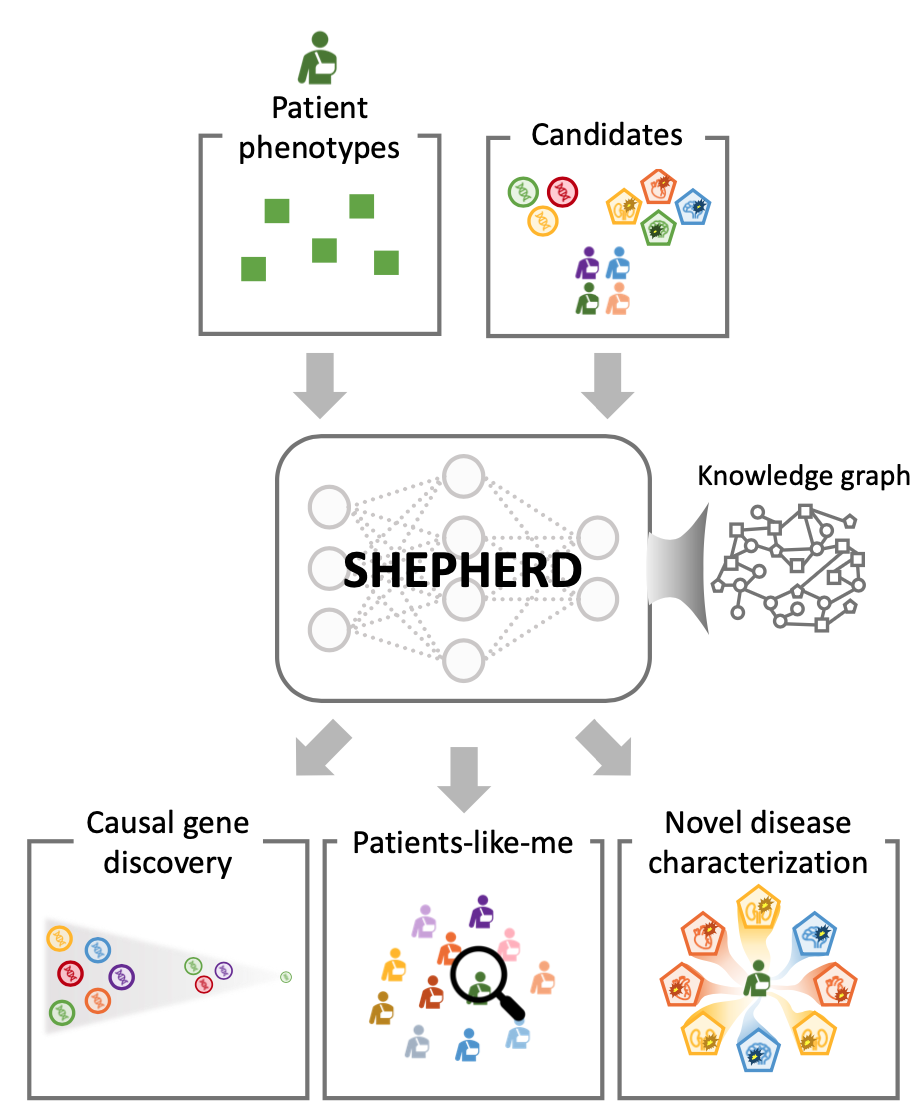
Overview of SHEPHERD
SHEPHERD is first deep learning approach for individualized diagnosis of rare genetic diseases. It provides multi-faceted diagnosis of patients with rare genetic conditions. SHEPHERD operates at multiple points throughout the rare disease diagnosis process to perform causal gene discovery, retrieve “patients-like-me” with similar conditions, and provide interpretable names for novel disease presentations.
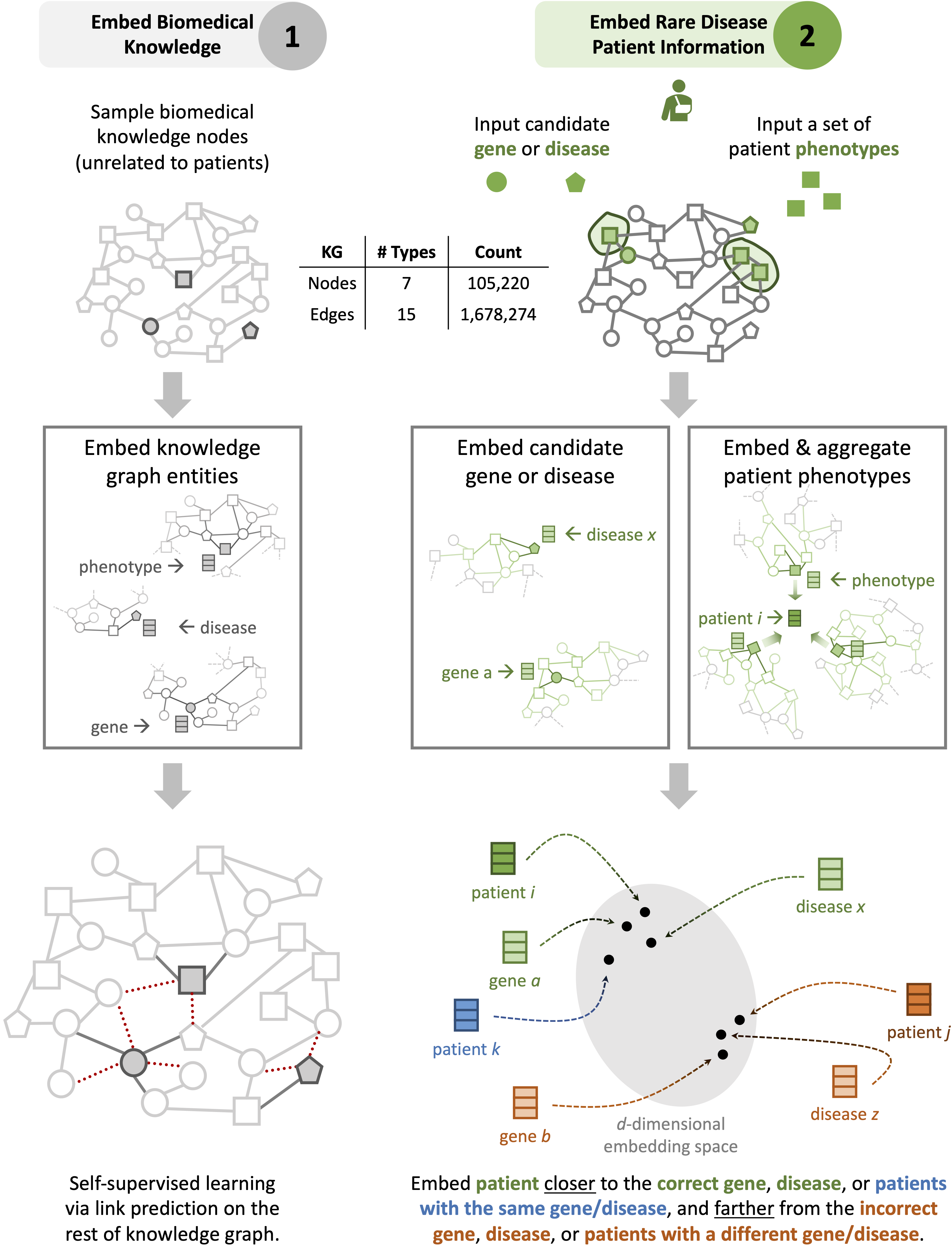
To overcome the limitations of supervised AI, SHEPHERD performs label-efficient training via knowledge-grounded metric learning, a deep learning paradigm uniquely suited for diagnostic analyses where patient diagnostic labels are incredibly scarce. This capability is achieved by projecting patient phenotypes in an embedding space whose geometry is optimized by the broader knowledge of phenotypes and genes.
When a new patient arrives, SHEPHERD projects the patient into the embedding space such that the patient is close to their most promising causal gene and disease and other patients with the same gene or disease. Using this embedding space for prediction, SHEPHERD nominates a patient’s genes and diseases even when no other patients are diagnosed with the same disease.
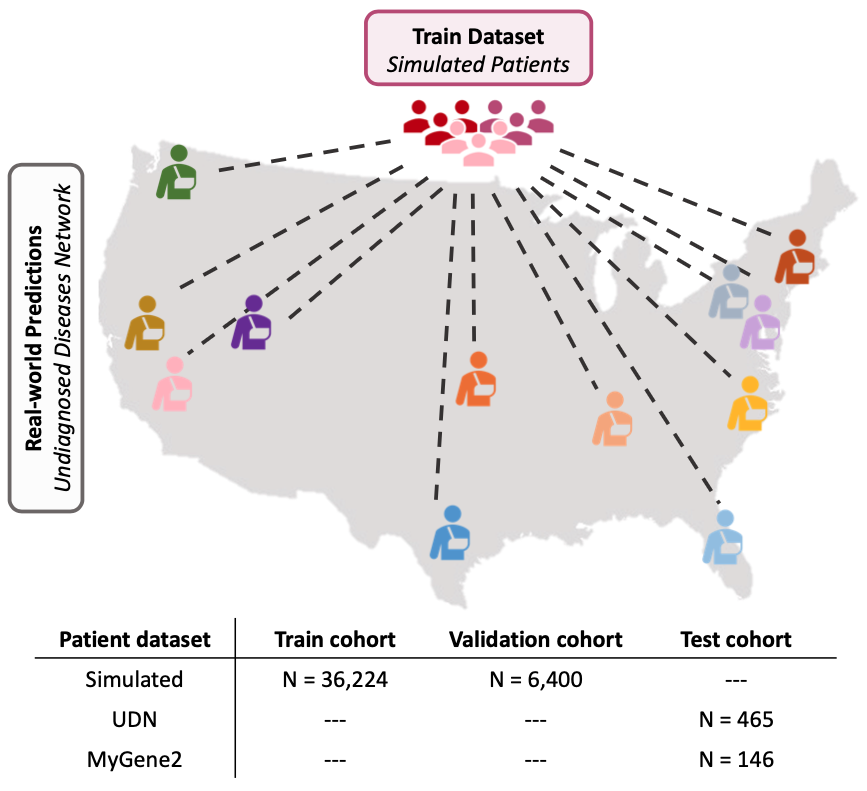
Evaluation on external patient cohorts
We evaluate SHEPHERD on an external cohort of patients in the Undiagnosed Diseases Network (UDN), a nationwide US study with 12 clinical sites that diagnose patients with rare, difficult to diagnose genetic conditions. SHEPHERD identifies the correct causal gene in 40% of patients spanning 16 disease areas.
Further, we show that SHEPHERD produces models with strong predictive performance across disease areas, clinical sites, and diagnosis years, successfully mitigating data mismatches between patient groups and pointing to SHEPHERD’s ability to minimize model performance gaps between patient groups.
SHEPHERD also generates meaningful patient representations that capture patient similarity and enable retrieval of “patients-like-me” with similar genetic conditions. Finally, our analyses demonstrate that SHEPHERD can nominate the correct diagnosis for patients with novel genetic diseases and provide interpretable characterizations of novel disease presentations.
Publication
Few Shot Learning for Phenotype-Driven Diagnosis of Patients with Rare Genetic Diseases
Emily Alsentzer*, Michelle M. Li*, Shilpa N. Kobren, Ayush Noori, Undiagnosed Diseases Network, Isaac S. Kohane and Marinka Zitnik
npj Digital Medicine 2025 [medRxiv]
* Equal Contribution
@article{alsentzer2025few,
title={Few Shot Learning for Phenotype-Driven Diagnosis of Patients with Rare Genetic Diseases},
author={Alsentzer, Emily and Li, Michelle M and Kobren, Shilpa N and Noori, Ayush and Undiagnosed Diseases Network and Kohane, Isaac S and Zitnik, Marinka},
journal={npj Digital Medicine},
url={https://www.nature.com/articles/s41746-025-01749-1},
year={2025},
publisher={Cold Spring Harbor Laboratory Press}
}
Data availability
Datasets are available for download via Harvard Dataverse. To maintain the directory structure while downloading the files, make sure to select all files and download in the original format. We provide the following datasets for training SHEPHERD:
- Rare disease knowledge graph
- Disease-split train and validation sets for simulated patients
- A cohort of MyGene2 patients
While the UDN dataset cannot be released in its entirety due to privacy concerns, anonymized UDN data has been deposited in dbGaP (accession phs001232) and PhenomeCentral. Phenotypes and causal variants and genes related to UDN diagnoses are shared publicly in ClinVar.
Code availability
Pytorch implementation of SHEPHERD is available in the GitHub repository.
We also provide an interactive demo to explore SHEPHERD’s predictions through a visual interface in the HuggingFace Space.