Protein structure is central to biological function. Enabling structure-native foundation models requires representations that discretize continuous backbone geometry while preserving global consistency and multi-scale organization. Existing protein structure tokenizers rely on fixed-size codebooks or continuous latent vectors, limiting interpretability, resolution control, and transfer across architectures.
Here we introduce GeoBPE, a geometry-grounded protein structure tokenizer inspired by byte-pair encoding. GeoBPE transforms continuous backbone conformations into discrete, hierarchical “sentences” of structural motifs. At each step, GeoBPE identifies frequent motif pairs, clusters them via k-medoids to form representative prototypes, and replaces occurrences with learned geometric primitives. To prevent geometric drift introduced by local quantization, GeoBPE performs differentiable inverse kinematics to optimize boundary glue angles under an SE(3) end-frame loss, preserving global fold integrity.
We evaluate GeoBPE on large-scale structural datasets and functional tasks. GeoBPE achieves strong compression–distortion tradeoffs, robust out-of-distribution generalization, and data-efficient training. It improves representation learning across binding site prediction, fold classification, and structural property prediction. Tokens align with CATH functional families and support expert-interpretable case studies. GeoBPE establishes a principled foundation for structure-native protein language models.
Overview of GeoBPE
Protein language models trained on sequence data capture evolutionary constraints but do not explicitly encode backbone geometry. Modeling structure directly requires discretizing continuous 3D conformations into symbolic units that preserve fold-level consistency and functional organization. The core question is how to transform noisy, multi-scale backbone geometry into discrete tokens without sacrificing global structure.
An effective protein structure tokenizer should satisfy three properties:
- Hierarchical vocabulary: Learn reusable structural motifs that compose into higher-order fold elements.
- Geometric fidelity: Preserve global SE(3) consistency after local quantization.
- Multi-resolution control: Allow adjustable compression–distortion tradeoffs across tasks.
Existing approaches based on vector quantization use fixed-size codebooks and latent embeddings. These representations lack hierarchical structure and provide limited control over resolution.
GeoBPE addresses this gap by extending byte-pair encoding to continuous protein geometry. It alternates between local motif merging and global geometric correction.
-
Motif discovery: GeoBPE identifies frequent adjacent motif pairs (Geo-Pairs), clusters their occurrences via k-medoids, and introduces representative geometric prototypes into a growing vocabulary.
-
Hard quantization: Each occurrence is replaced with its nearest medoid prototype, producing discrete motif tokens.
-
Glue-aware refinement: Quantization introduces geometric drift. GeoBPE corrects this by optimizing boundary glue angles using differentiable inverse kinematics under an SE(3) end-frame loss.
The result is a hierarchical merge tree that segments a backbone into multi-scale structural motifs. The learned vocabulary provides an interpretable representation of protein structure.
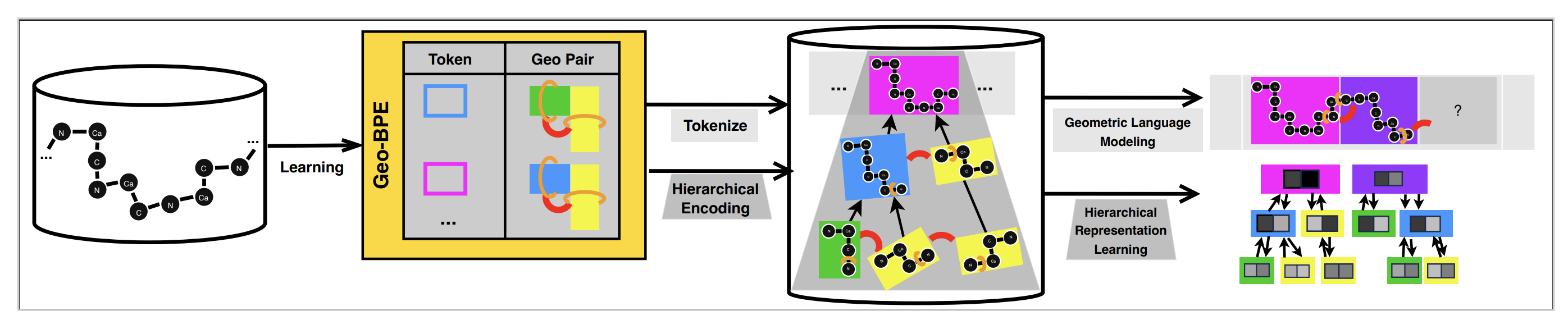
GeoBPE: Geometry-Grounded Byte-Pair Encoding
GeoBPE builds a discrete structural alphabet while preserving fold-level consistency. Given backbone coordinates, it proceeds in iterative merge steps.
GeoBPE begins at the residue level. Each residue is clustered via RMSD into representative prototypes, forming the initial vocabulary. The backbone is rewritten using these residue-level motifs.
At each iteration:
- The most frequent Geo-Pair is identified.
- All occurrences are gathered across the dataset.
- K-medoids clustering produces representative prototypes.
- Occurrences are replaced by their assigned prototypes.
- The merge hierarchy is updated.
This process mirrors byte-pair encoding but operates in geometric space rather than symbol space. Vocabulary size and resolution are controlled through the number of medoids and merge iterations.
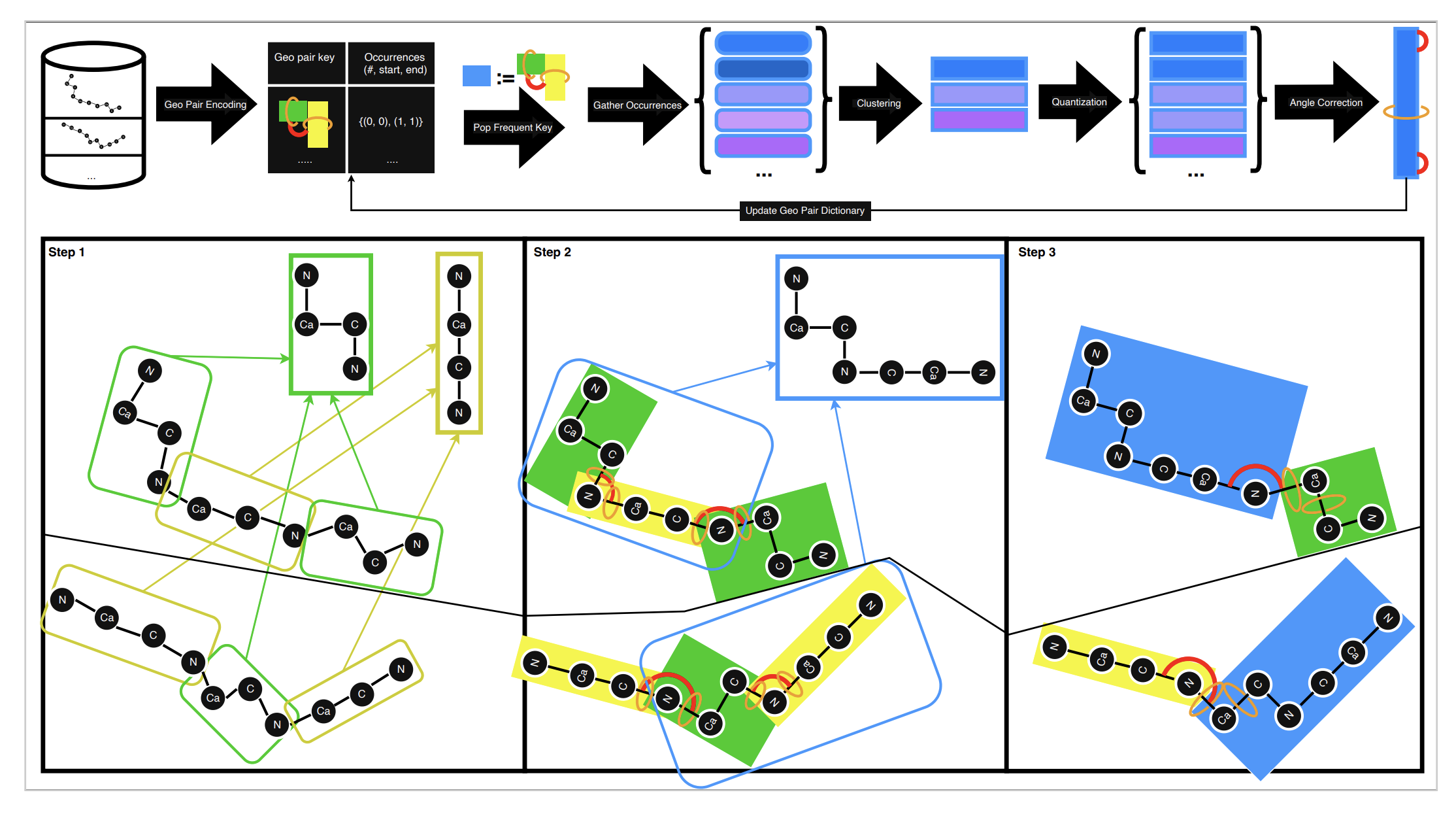
Replacing motif pairs with medoid prototypes introduces geometric drift. Without correction, accumulated local errors distort the global fold. GeoBPE addresses this through glue-aware refinement.
Each motif boundary is parameterized by three glue angles. After quantization, GeoBPE optimizes these angles via differentiable forward kinematics to minimize an SE(3) end-frame loss between reconstructed and original structures.
This step:
- Preserves global backbone consistency.
- Prevents drift accumulation across merges.
- Enables stable multi-step hierarchical decomposition.
Rigid-body refinement is essential. Removing glue optimization substantially increases RMSD and degrades fold integrity.
Multi-Resolution and Architecture-Agnostic Protein Tokenizer
GeoBPE supports adjustable resolution by construction. As the vocabulary grows, GeoBPE captures a larger fraction of backbone variability and improves reconstruction fidelity. With fewer merges and a smaller vocabulary, it produces coarser motifs that favor abstraction and representation learning. This makes resolution controllable, allowing the same tokenizer to serve compression, downstream transfer, or structure language modeling depending on the target use case.
GeoBPE is also architecture agnostic. It outputs a hierarchical merge tree that can coarsen residue-level embeddings from large protein language models into motif-level and protein-level representations, and it can be paired with a transformer to model structure tokens and generate backbones by language modeling.
Because GeoBPE tokens are observed structural medoids, they remain interpretable rather than opaque latent vectors. The resulting motifs align with functional domain boundaries and recurrent structural patterns.
Publication
Protein Structure Tokenization via Geometric Byte Pair Encoding
Michael Sun, Weize Yuan, Gang Liu, Wojciech Matusik, Marinka Zitnik
International Conference on Learning Representations, ICLR 2026
@article{sun2026protein,
title={Protein Structure Tokenization via Geometric Byte Pair Encoding},
author={Sun, Michael and Yuan, Weize and Liu, Gang and Matusik, Wojciech and Zitnik, Marinka},
journal={International Conference on Learning Representations, ICLR},
year={2026}
}
Code and Data Availability
Pytorch implementation of GeoBPE is available in the GitHub repository.